I Transferred Data Between Perplexity, Google and Claude with a Text File. Healthcare Still Faxes.
Can We Solve the Interoperability Problem with Markdown Files?
On the same afternoon I was doing two things that I realized were related but oddly at odds with each other.
First, I was building a research-to-execution pipeline across three AI tools. I used Perplexity to pull validated literature, exporting a clean markdown document. That document went into a NotebookLM notebook I’ve curated with prompt engineering best practices. I used NotebookLM as a metaprompt to synthesize a precise prompt guided by the research. That prompt went into Claude to execute the task. The whole thing ran on plain language. There were no API calls, database schemas or IT tickets. The exchanges between platforms were text passing between systems, but each was structured in a way that the receiving tool was able to easily read it and leverage each tool’s unique strengths to maximize the workflow.
Later, I worked on an Epic related project where we were tracking quality documentation attached to a particular type of text. We tagged a discrete Epic SmartData Element to a note called a SmartText. The note had nuance about the details of documentation, in this case, reevaluation of a patient’s condition after a sepsis intervention. We use this technique quite frequently when we capture quality metrics using Epic documentation tools. The Smart Data Element captured the completion of the documentation as a discrete value but lost all the nuance.
I realized that I was addressing similar interoperability problems in different ways. One workflow was elegant. The other was clunky.
Why We Built What We Built
The promise of healthcare data interoperability was never really about moving files. It was about making clinical information computable and usable. A system at one hospital receiving data from another shouldn’t just receive it. It should be able to do something with it like risk stratify a population, surface a drug allergy or close a care gap.
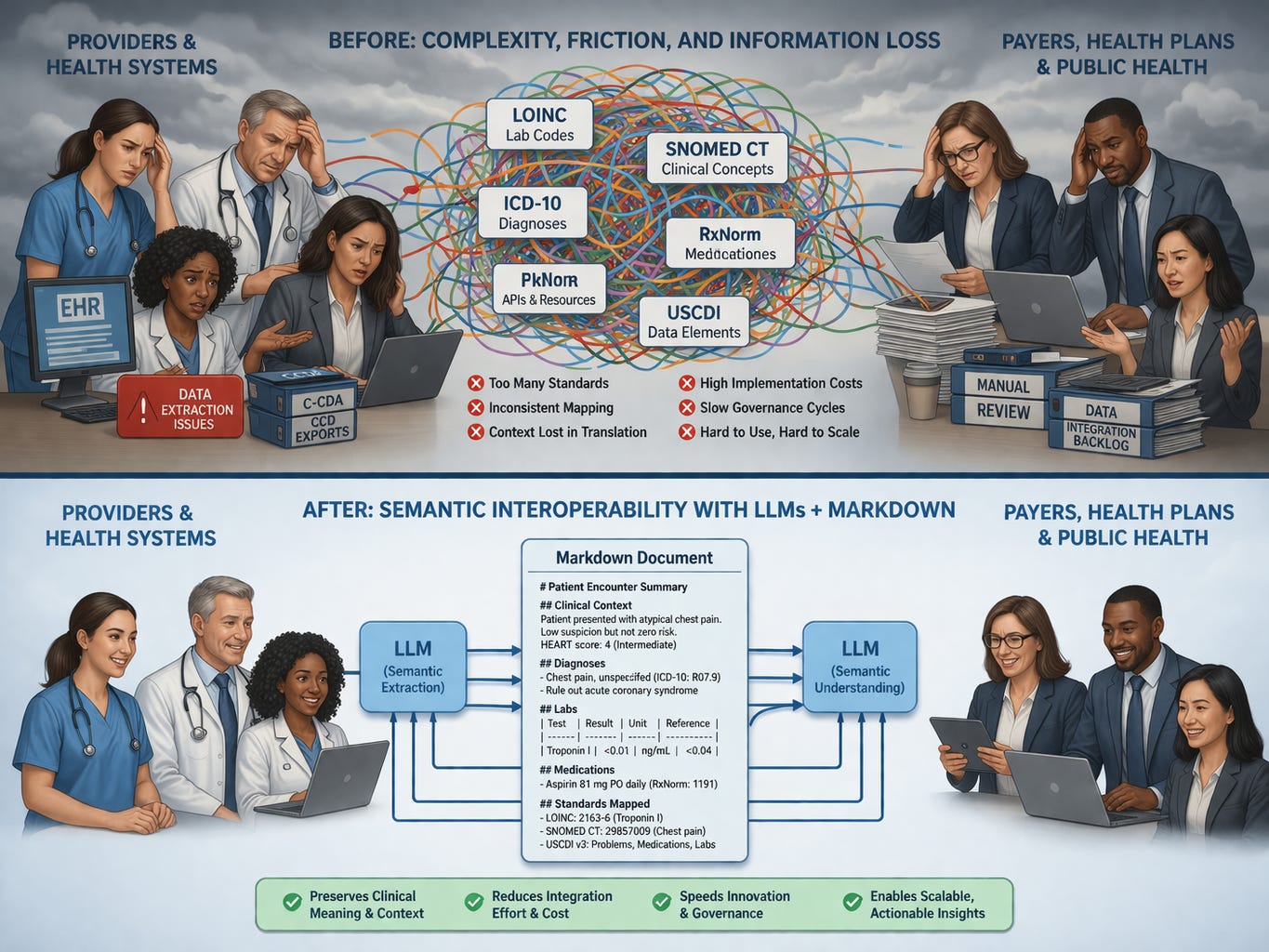
To accomplish that, we have traditionally required reams of structured data fields to be machine-readable. So we built standards like HL7 v2 for messaging, C-CDA for documents, and FHIR for APIs. Underneath all of them, we had an alphabet soup of terminology standards: LOINC for labs, SNOMED CT for clinical concepts, ICD for diagnoses, CPT for procedures. We created USCDI to define the minimum dataset that should be exchangeable at all.
These were and are not bad ideas. LOINC alone represents decades of work by people who cared deeply about making lab results mean the same thing across institutions. The FHIR specification enabled a generation of patient-facing apps that wouldn’t exist otherwise.
But they require every piece of clinical meaning to be pre-enumerated, defined and locally configured in advance. On top of this the underlying controlled vocabulary is governed typically on a cycle measured in years or decades, not weeks.
That requirement creates two simultaneous failures. They’re related but distinct, and conflating them is part of why the solutions keep missing.
Failure One: Clinical Meaning Doesn’t Compress
When you force clinical information into a controlled vocabulary, you are not translating it. You are compressing it. Compression always loses something.
A LOINC code for a hemoglobin A1c tells a receiving system that a certain test was performed and a certain value returned. It does not tell the system that the ordering physician had already discussed with this patient that the result, given their recent illness and the fact that they’d been unable to afford insulin for three weeks, should be interpreted differently than the raw number suggests. That context doesn’t have a tidy code. The single result is also just a small part of the package. Attached to it are all sorts of metadata such as date, resulting location and reference values. These create a tsunami of data when we might have heuristically been expecting a trickle. (See my previous piece on this.)
SNOMED CT has over 350,000 active concepts. It still cannot neatly capture what a clinician means by “atypical chest pain, low suspicion but not zero risk” on a patient whose HEART score is intermediate. We are forced to remove much of our cognitive clinical reasoning into a rigid code.
This is not a flaw in the standards. It is a structural feature of what standards are. A controlled vocabulary is, by definition, a closed set of pre-agreed meanings. Clinical reality does not neatly fit into a closed set.
So we make compromises. We find the nearest code. We tag the closest concept. We attach the SmartData Element that fulfills a quality reporting metric, but does not transmit the underlying meaning of what we actually documented.
On the other hand, when you don’t compress, you get a mess. CCD documents were one of the more prominent efforts to address interoperability early in the EHR frenzy. It seemed like a good idea: send a comprehensive summary to a stakeholder like a primary care physician after an encounter. However, there was a big wrinkle. As a CMIO, I fielded multiple complaints for years about CCDs arriving by fax. The documents ran dozens, sometimes hundreds of pages. I understood the irony every time. The CCD was never designed to be printed and fed through a phone line. It was built to be ingested by a system, parsed, and turned into something actionable. But I could not point to a single organization that had actually done that. The CCD arrived, it got filed, and the clinician went looking for the relevant paragraph the same way they would have searched through a paper chart. I knew the proper fix: configure systems to ingest and parse these documents computationally. I also knew that doing it properly would require a substantial configuration project nobody had the budget or appetite for. So I kept faxing.
Failure Two: The Governance Architecture Is the Problem
Semantic compression would be more tolerable if the standards could evolve quickly. They cannot.
Adding a concept to SNOMED CT requires a formal proposal, a review cycle, balloting, version management, and downstream updates to every implementation guide that references it. USCDI updates happen annually at best. FHIR implementation profiles take years to reach consensus and broader adoption. By the time a governance body ratifies a new data element for, say, social determinants of health screening results, the clinical landscape has already moved on.
COVID was an example of the lag and friction we have in governing terminology sets. When COVID arrived in early 2020, it did not have a diagnosis code. ICD-10 had no entry for a disease that had not existed when the vocabulary was last updated. For several weeks, clinicians documenting patients who almost certainly had COVID were coding them as upper respiratory infection, viral bronchitis, or influenza — whatever was nearest and defensible. The clinical picture was clear. The patients had COVID. But the machine readable data was muddy at best because we had no code. Surveillance systems, population health dashboards, and epidemiological tracking were all running on a dataset that was systematically mislabeling the most important diagnosis in a generation. We were able to correct this within weeks but that was only because a once in a century pandemic tends to cause things to move quickly. For more mundane situations like a chest pain patient at low-intermediate risk, that label will likely never come. The subtlety will only be visible in the unstructured note.
That is not a failure of implementation. That is the structural ceiling of any system that requires clinical reality to wait for governance approval before it can be represented in data. Every new interoperability connection requires a mapping exercise. Source field to target field. Source vocabulary to target vocabulary. And because no two organizations implement the same standard identically, because FHIR conformance does not guarantee interoperability in practice, every connection becomes a bespoke translation project. Every translation project requires analysts. Analysts require time, budget, and organizational will.
The governance overhead is not incidental to the standards. It is the price of the deterministic model. When you require that every meaning be pre-agreed and pre-enumerated, you require a process for agreeing and enumerating. That process is slow because the domain is complex and the stakeholders are many and decision makers have to accommodate multiple downstream purposes for the terminology. The semantics are only part of it. Regulatory compliance, quality reporting, billing integrity, and any number of other downstream uses all have claims on how a concept gets defined.
Here is the part that matters for anyone working in policy: this architecture doesn’t just move slowly. It structurally disadvantages any clinical concept that doesn’t already have a constituency large or significant enough to push it through governance. Discrete tags for rare conditions, novel workflows, emerging social determinants, or subtle labeling of gray areas are addressed over decades, if they are addressed at all.
What I Was Actually Doing with Those Markdown Files
Let’s go back to the pipeline I described.
Perplexity doesn’t export to a schema. It exports to prose called “markdown.” It is text but is clean, structured, and machine readable by LLMs. NotebookLM doesn’t parse fields. It reads the document and synthesizes from it because the markdown formatting makes this easy. Claude doesn’t look up a code table. It interprets language in context.
At no point in that workflow did I define a data dictionary. I didn’t map fields. I didn’t register a terminology server. I didn’t create a spreadsheet or database. I wrote what I needed, structured it loosely with markdown headers and bullets, and passed it forward. Each system interpreted it correctly for its purpose because each system was capable of reading meaning, not just parsing structure.
Before going further, let me give a quick orientation for anyone who hasn’t worked directly with these tools.
Markdown is plain text with lightweight formatting conventions. A pound sign makes a heading. A dash makes a bullet. Double asterisks around a word make it bold. If you have ever typed an asterisk around a word in a text message for emphasis, you have used markdown instinctively. It reads cleanly as plain text to any human. It renders as formatted content in almost every modern application that can read TXT files. You do not need special software to create it, read it, or edit it. A clinician could write it. A policy analyst could write it.
LLMs were trained on the internet, and the internet is largely written in markdown. When you pass a well-structured markdown document to an LLM, it does not need a schema to interpret it. It reads it the way a well-read person reads a structured memo: extracting what is relevant, inferring relationships, understanding context from proximity and hierarchy.
That is a meaningful contrast with the current healthcare interoperability model. XML schemas, HL7 segments, FHIR resource definitions — these require a parser that has been explicitly programmed to expect a specific structure at a specific location. If the structure deviates, the parser fails or silently drops data. A language model reading markdown does not fail when a field is missing or phrased differently. It interprets what is there and works with it.
The difference is what’s consuming it: a language model capable of extracting whatever is relevant for the task at hand, rather than a parser that can only recognize what it was explicitly programmed to look for.
A decade ago, that kind of multi-system integration required explicit programming, API documentation, agreed schemas, and developers who understood each system’s internals. Today it can run on plain language because the systems on both ends understand plain language if you use language models. The integration layer is no longer a rigid contract between machines. It is a flexible interpretation between models capable of reading context.
Now extend that concept to healthcare interoperability.
What an LLM-Native Exchange Layer Could Look Like
My proposal is not to replace standards with chaos. It is to add a layer between clinical documentation and downstream consumption that uses language models to do what language models are actually good at: reading context, extracting meaning, and simultaneously produce structured output tailored to a specific use case.
At each transition point — a patient transferred from one system to another, a referral sent across organizations, a population health query running against records from multiple sources — an LLM can generate a markdown document. Each document is use-case-specific. The ED physician receiving a transfer gets a packet emphasizing active problems, medications, allergies, and recent interventions pertinent to the ED scenario. The care manager gets social determinants, care gaps, and recent utilization. The quality analyst gets the discrete elements relevant to the measure being run.
The same underlying source data can create different markdown documents. Each workflow would have different prompts producing different outputs. Anyone who has received and read healthcare documentation understands the phenomenon of simultaneously sparse and verbose documentation: none of the content you need is there but there are large volumes of information you don’t need. The markdown approach avoids the pre-enumeration or rigid structures that causes this odd contradiction.
When the receiving organization ingests a curated markdown document, their own LLM processes it into whatever internal structure they need. The translation happens in language, which preserves context, rather than in code mapping, which compresses it. It is the solution for the CCD that we never quite accomplished.
This is not free of work. Prompts need to be designed and validated. Quality checks need to be built. Edge cases will surface. But the work is different in character from traditional interoperability work. It does not require a governance body to ratify a new field. It does not require a bilateral mapping agreement between analysts at two organizations who will both be gone in eighteen months. It requires good prompt engineering and a local validation pipeline. These are far more flexible and accessible than waiting for ICD 11.
Consider what just happened with CMS-0053. After nearly thirty years of failed attempts, HHS finally finalized a standard for electronic claims attachments, the supporting clinical documentation payers request when adjudicating a claim. The standard it adopted: X12 275 plus HL7 C-CDA templates. It is necessary and long overdue. But it is also a clear illustration of what the deterministic model costs in practice. Each sending organization must configure their system to generate conformant C-CDA documents. Each receiving payer must configure their system to parse them. Every edge case, such as a field populated differently, a template implemented loosely, or a document that is technically conformant but structurally idiosyncratic, becomes a configuration problem that lands on an analyst’s desk that goes into a Service Now backlog.
Now consider the alternative. A sending organization writes a prompt that pulls the relevant clinical context from the patient record and produces a structured markdown document. The receiving organization runs that document through their own prompt to extract the specific elements needed for their adjudication workflow. The sending prompt is written by someone who understands the clinical and operational context. The receiving prompt is written by someone who understands the payer’s workflow. There is no schema negotiation. There is no conformance testing against a ballot-voted implementation guide. The implementation can be measured in days or weeks instead of months to years.
Brendan Keeler framed this well: fax achieves two things simultaneously that every proposed electronic alternative has failed to match — equivalent ubiquity: it works because every counterparty already has it, and equivalent workflow flexibility: you can fax anything to anyone and rely on a human receiver to interpret whatever arrives, regardless of format. Every electronic standard has solved the format problem while failing the ubiquity problem, because the configuration burden is high enough that smaller organizations, rural providers, and under-resourced practices get left behind.
An LLM-mediated markdown exchange also can achieve both. Any organization that can access a language model and write a prompt has the infrastructure. Any clinical content that can be described in language (ie: everything) can be exchanged. The interoperability layer is no longer a rigid technical contract between two systems. It is a flexible interpretation between two models, each operating in the context that matters to them.
Think of it as USCDI providing the floor: the minimum discrete data that travels as structured fields. Everything above that floor, like the contextual and semantically complex content that LOINC and SNOMED were never designed to carry, travels as a use-case-specific markdown document interpreted by models on both ends.
The Hallucination Objection
Of course, people will worry because LLMs hallucinate. You need deterministic infrastructure because you cannot build patient safety and economic models on a probabilistic text model.
That is a fair concern, but it is the wrong comparison.
Look at the problem list on any moderately complex patient moving through a real health system. You will find diagnoses that resolved years ago still listed as active. Medications are duplicated across transitions of care and also list items that are no longer valid. Allergies are documented twice with different terminology. Procedures are coded to the nearest available code rather than the actual procedure performed. Conditions are carried forward from a note that carried them forward from a note written by a resident who wasn’t certain and didn’t want to delete anything.
Our current infrastructure produces noise at a rate that would concern anyone analyzing it systematically. We just don’t have a high-profile, trendy word for it like “hallucinations.” They are human transcription errors, workflow shortcuts, semantic compression, and copy-forward documentation that compound in ways that have never been cleanly benchmarked against a ground truth. Realistically, we don’t even know what the ground truth is. The difference is that we’re accustomed to it, we’ve built workflows around it, and nobody is publishing error rates.
The question is not whether an LLM-based exchange layer introduces error. Every step in our current system introduces error. The question is whether it introduces more error than the status quo, of what kind, and whether those errors are more or less recoverable than the ones we already accept. That is an empirical question worth taking seriously. Reflexively dismissing LLM-based approaches because of hallucination risk, as though the current system is delivering clean ground truth, is not a serious position.
What the Standards Are Still For
None of this argues for abandoning FHIR, retiring LOINC, or walking away from USCDI. It argues for using them where they belong.
LOINC codes still work as stable identifiers, references that tell a receiving system what kind of thing it is looking at before the language model interprets it. FHIR still provides a transport framework that ensures basic structural compatibility at the API layer. USCDI still defines a useful floor for what data should be exchangeable. These are scaffolding.
The argument is about what happens in the gaps. The contextual, the nuanced, the semantically complex content are areas that controlled vocabularies were never designed to carry. That is where language models are capable and where the current approach is failing. The standards don’t need to be replaced. They need to be relieved of work they were never suited to do.
The Distance That Matters
We spent thirty years trying to make clinical meaning machine-readable by forcing it into structures that machines could parse with 1990s technology. We built governance architectures to manage those structures that now move slower than the clinical problems they’re supposed to represent.
We have different technology now. Technology that reads language, holds context across a document, and extracts what is relevant for a specific purpose. Technology that makes errors, yes, as does every physician who has shrugged and documented a “close enough” ICD code.
The markdown document passing between my tools that afternoon wasn’t a workaround. It was a working demonstration of what interoperability looks like when the systems on both ends are capable of understanding meaning rather than just parsing structure.
We built the old model because it was the only model available. It is not the only model anymore.
John Lee is an emergency physician and Epic consultant who helps health systems bridge the gap between Epic’s capabilities and operational reality. He specializes in data architecture, registry optimization, and making Epic’s tools actually deliver results.
If you need help configuring your Epic environment to support these capabilities, connect with him on LinkedIn or via his website.


See, THIS is the actual use case for AI/LLMs in medicine (now that we're starting to get a handle on ambient AI-assisted documentation to make doctor's hate their jobs less, that is). Also very clearly written by a pragmatic EM doc. Thanks so much for this excellent read!
- A (hopefully) pragmatic EM doc
I very much enjoyed this one John. Well done